Article Title :
Distribution and Trend Analysis of COVID-19 in India: Geospatial Approach 
Murugesan Bagyaraj
,
SHANKAR KARUPPANNAN
 ,
Alemayehu Tenaw Mengistie
,
Gnanachandrasamy Gopalakrishnan
,
Alemayehu Tenaw Mengistie
,
Gnanachandrasamy Gopalakrishnan
4 (2020)
1-9
Spatial Distribution , India , GIS , COVID-19 , Coronavirus


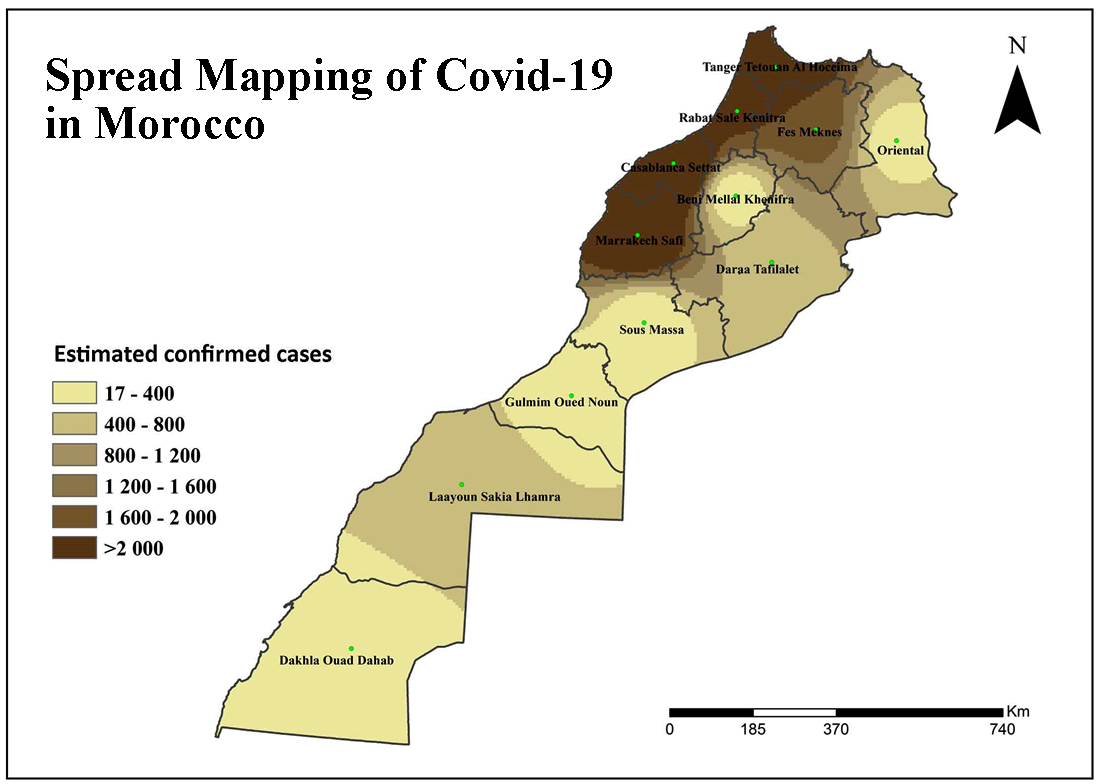
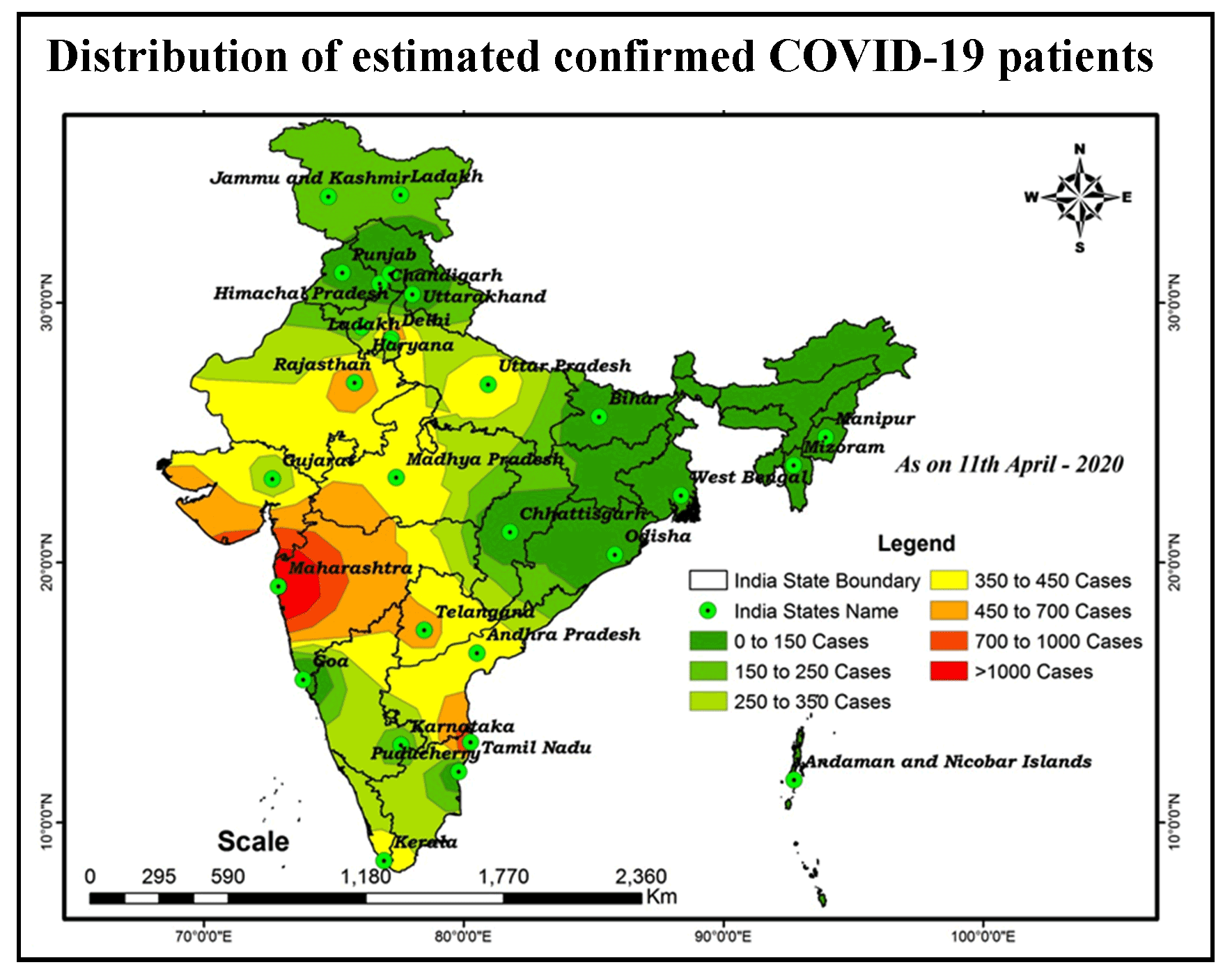
COVID-19 Coronavirus is now one of the most contagious diseases of the recently discovered and spread across the China in 2019 and has received global attention. In most COVID-19-infected individuals, respiratory symptoms should be mild to moderate and improve without the need for medical care. The risk of serious disease is higher for senior citizens and people with serious health problems, such as heart disease, diabetes, severe respiratory disease, and cancer. The World Health Organization (WHO) has formally declared the outbreak of COVID-19 to be a global pandemic. As on 11th April 2020 in India the largest number of persons testing positive for COVID-19 since the outbreak earlier month with samples of people, mostly contacts of already confirmed patients, rendering positive. In India total confirmed cases 7364, 633 are cured/discharged, with 240 deaths had been reported by the Ministry of Health and Family Welfare Government of India. The aim of the research is to analyze the spatial distribution of COVID-19 and its trends with the help of GIS software. At this time, there are no precise antibiotics or treatment options for COVID-19. Besides, several ongoing clinical studies are assessing effective treatments. The best way to protect and sluggish transmission should be well advised about the current COVID-19 virus, the disease it triggers and also how it continues to spread. Therefore, monitoring active ties using GIS spatial analysis is very important to control such as a COVID-19 virus spreading problem.
COVID-19 is an increasingly pandemic disease which haunts social life on a daily basis.
GIS can support the fight against infectious diseases caused by outbreaks and epidemics of COVID-19.
Distribution patterns of disease transmission are illustrated by the GIS tool and predict future confirmed cases.
Distribution and trend analysis shows support for decision-making in the transmission and prevention of epidemics.
This analysis may provide valuable information to support the government's monitoring and forecasting of the spread of the virus across small and large areas.
Spatial analysis may be used to identify threats to disease, patterns of time and space epidemics and hotspot infections.
Chan, J. F., Yuan, S., Kok, K. H., To, K. K., Chu, H., Yang, J., Xing, F., Liu, J., Yip, C. C.-Y., Poon, R.W.-S., Tsoi, H.W., Lo, S. K.-F., Chan, K.H., Poon, V. K.-M., Chan, W.-M., Ip, J.D., Cai, J.P., Cheng, V.C.-C., Chen, H., Hui, C. K.-M. and Yuen, K. Y., 2020. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating personto-person transmission: A study of a family cluster. Lancet, 395(10223), 514-23.
Childs, C., 2004. Interpolating surfaces in ArcGIS spatial analyst. ArcUser, July-September, 3235, 569.
Guan, W. J., Ni, Z. Y., Hu, Y., Liang, W. H., Ou, C. Q., He, J. X., Liu, L., Shan, H., Lei, C.-L., Hui, D. S. C., Du, B., Li, L. -J., Zeng, G., Yuen, K.-Y., Chen, R., Tang, C., Wang, T., Chen, P., Xiang, J., Li, S., Wang, J., Liang, Z., Peng, Y., Wei, L., Liu, Y., Hu, Y., Peng, P., Wang, J., Liu, J., Chen, Z., Li, G., Zheng, Z., Qiu, S., Luo, J., Ye, C., Zhu, S., and Zhong, N., 2020. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 1-13.
Kang, C. K, Song, K. H, Choe, P. G, Park, W. B, Bang, J. H, Kim, E. S, Park, S. W., Kim, H. B., Kim, N. J., Cho, S., Lee, J.-K., and Oh, M., 2017. Clinical and epidemiologic characteristics of spreaders of middle east respiratory syndrome coronavirus during the 2015 outbreak in Korea. J Korean Med Sci. 32(5), 744-9.
Lu, R., Zhao, X., Li, J., Niu, P., Yang, B., Wu, H., Wang, W., Song, H., Huang, B., Zhu, N., Bi, Y., Ma, X., Zhan, F., Wang, L., Hu, T., Zhou, H., Hu, Z., Zhou, W., Zhao, L., Chen, J., Meng, Y., Wang, J., Lin, Y., Yuan, J., Xie, Z., Ma, J., Liu, W. J., Wang, D., Xu, W., Holmes, E. C., Gao, G. F., Wu, G., Chen, W., Shi, W. and Tan, W., 2020. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet, 395 (10224), 565-74.
Wu, F., Zhao, S., Yu, B., Chen, Y.M., Wang, W., Song, Z.G., Hu, Y., Tao, Z.-W., Tian, J. H., Pei, Y-Y.,Yuan, M. L., Zhang, Y. L., Dai, F.-H., Liu, Y., Wang, Q.-M., Zheng, J.-J., Xu, L., Holmes, E. C. and Zhang, Y.-Z., 2020. A new coronavirus associated with human respiratory disease in China. Nature. 579, 265-269.
Zhou, P., Yang, X.L., Wang, X.G., Hu, B., Zhang, L., Zhang, W., Si1, H._R., Zhu, Y., Li, B., Huang, C.-L., Chen, H.-D., Chen, J., Luo, Y., Guo, H., Jiang, R.-D., Liu, M. Q., Chen, Y., Shen, X.,-R., Wang, X., Zheng, X.-S., Zhao, K., Chen, Q.-J., Deng, F., Liu, L.-L., Yan, B., Zhan, F.-X., Wang, Y.-Y., Xiao, G.-F., and Shi, Z. -L., 2020. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 579, 270-273.